Main navigation
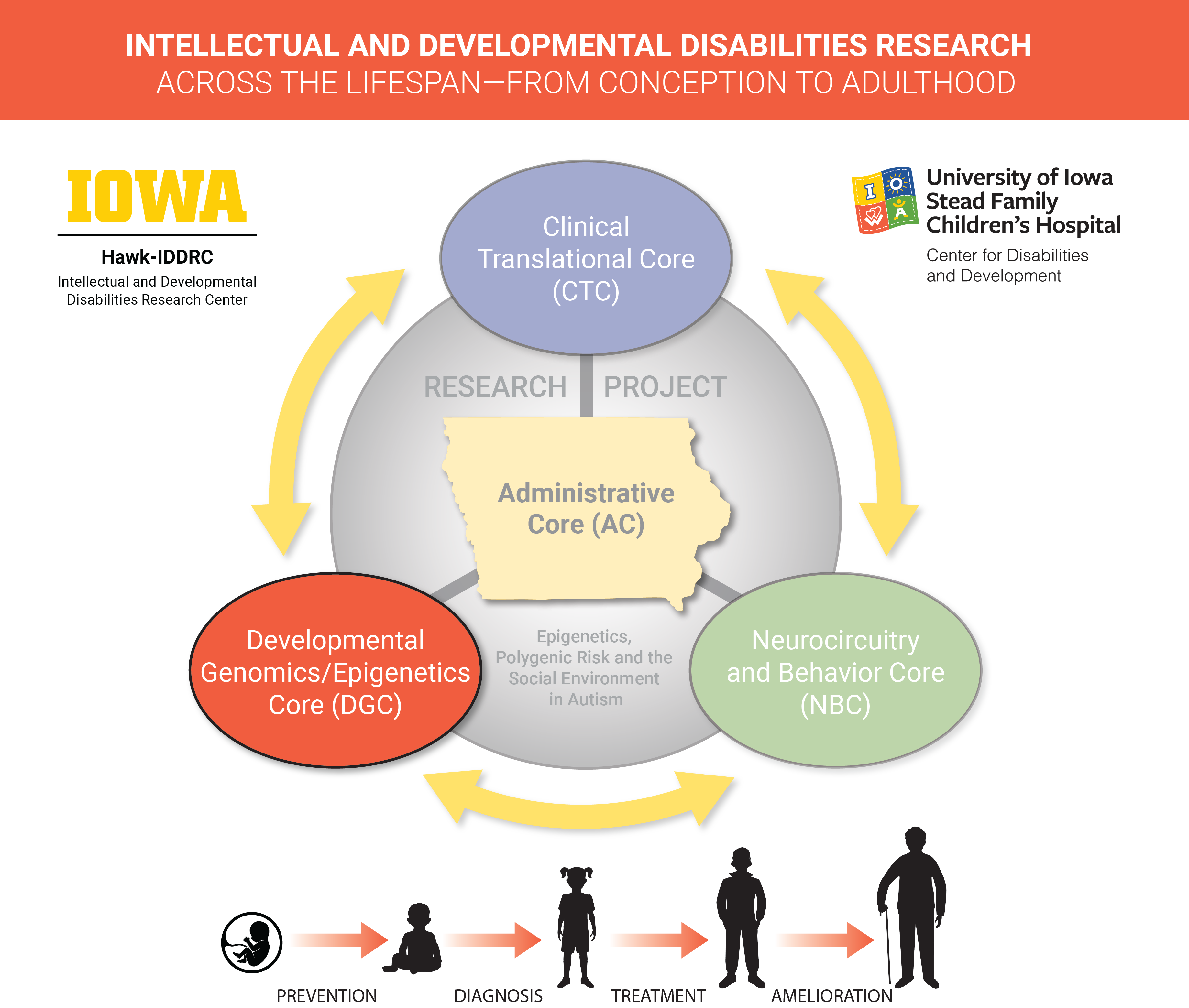
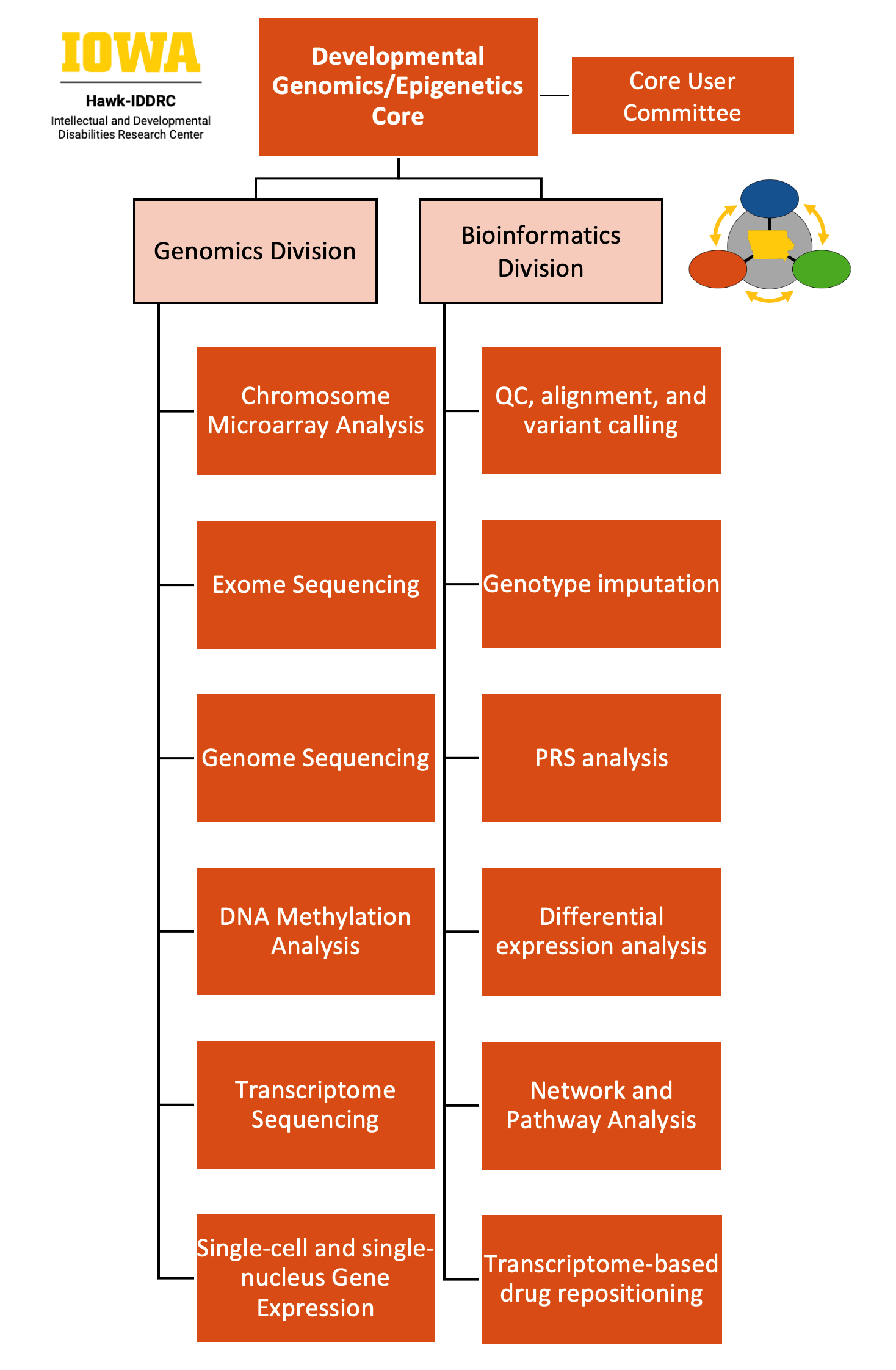
The Developmental Genomics/Epigenetics Core (DGC): The Developmental Genomics/Epigenetics Core supports innovative and cutting-edge research of IDDs across the lifespan – from conception to adulthood – and is tailored to a rural population. Under the leadership of Co-Directors Dr. Benjamin Darbro and Dr. Jake Michaelson, who together have substantial experience in experimental (Darbro) and computational (Michaelson) genomics, this Core provides broad expertise that encompasses model system development, high-throughput sequencing technologies (e.g., genome and exome sequencing, RNA sequencing, chromatin immunoprecipitation sequencing, methylation sequencing, and ribosome sequencing), and bioinformatics and computational analysis of the results. The Core is composed of (1) the Genomics Division, which provides high-throughput sequencing and array services including exome and genome sequencing, RNA-sequencing (including 10X Chromium single-cell sequencing), and array-based genotyping and methylation services under the direction of Dr. Benjamin Darbro, an accomplished genomics investigator; and (2) the Bioinformatics Division, which provides services for quality control, basic processing (e.g., alignment, variant calling, and expression quantification), and pipelines for calculating polygenic risk scores from data produced by array or sequencing services generated by the genomics division, and is led by Dr. Jake Michaelson, an experienced computational genomics investigator. Collectively, the available expertise and cutting-edge services provided by the DGC will catalyze IDD research by minimizing barriers for entry into genome-scale studies of neurodevelopment and cognitive function.